It's certainly no secret that the SARS-CoV-2 pandemic has changed everyone's lives in recent weeks. European countries are effectively closed, and people are more than ever being advised to self-isolate or work from home. It has put a tremendous amount of pressure on everyone.
Most of us at LinuxServer have now been mandated to work from home which has placed us in an entirely new working environment, closer to our families—primarily our spouses and children—providing us with new challenges! We are unable to dedicate as much time to volunteering on our support channels as we used to due to our days being turned upside-down. We don't all work in the IT industry either, with some of us in professions even more impacted by the pandemic. It's stressful. Hopefully those coming to us with queries on support for our images will understand that we may not be as responsive, or may be entirely unable to help until this thing blows over.
How you can help
Right now, scientists all over the planet are rushing to find new ways to protect us from a further outbreak of this virus. Using distributed computation, they are modelling various proteins in order to help them better understand how to create effective vaccines. This is where we all come in.
There are two large projects which allow people to run client programs on their own machines which share the computational workload of running complex tasks. Rather than having a single supercomputer crunching the numbers, tasks can be batched up and run simultaneously on thousands—or even millions—of machines at once. The great thing is we can install these clients on our home servers! And that's what we're asking you to do.
These projects are called Folding@home and BOINC, and are freely available for you to download and run on your own home machine. If you can spare the idle CPU on your NAS/PC/server, please consider running these clients to help speed up the process of finding an effective vaccine for COVID-19.
As always, we have provided a Docker image for both of these solutions which are outlined below. However, if you do not run Docker, each solution provides their own installer.
BOINC
By creating a shared infrastructure for computational tasks, BOINC allows multiple kinds of projects to utilise distributed computing:
BOINC lets you help cutting-edge science research using your computer (Windows, Mac, Linux) or Android device. BOINC downloads scientific computing jobs to your computer and runs them invisibly in the background. It's easy and safe. About 30 science projects use BOINC; examples include Einstein@Home, IBM World Community Grid, and SETI@home. These projects investigate diseases, study global warming, discover pulsars, and do many other types of scientific research.
As this is a multi-project solution, to specifically target the research of COVID-19, add the Rosetta@home project to your BOINC installation.
Installing
https://hub.docker.com/r/linuxserver/boinc
boinc:
image: linuxserver/boinc
container_name: boinc
environment:
- PUID=1000
- PGID=1000
- TZ=Europe/London
- GUAC_USER=boinc # Optional
- GUAC_PASS=<MD5SUM_OF_YOUR_PASSWORD> # Optional
volumes:
- /path/to/data:/config
ports:
- 8080:8080
devices:
- /dev/dri:/dev/dri #optional
restart: unless-stoppedConsidering this is a graphical tool, we are utilising our Apache Guacamole base image to give you graphical access to the client via a web browser. As optional layer of added security, you should provide the GUAC_USER and GUAC_PASS environment variables in your command. In this case the password provided should be a pre-hashed MD5 sum. Do not worry if this in particular is a bit above your head; feel free to continue without the authentication for Guacamole.
Access the web UI via http://<server_ip>:8080.
Once started, you will need to add the Rosetta@home project via
Tools -> Add Project:
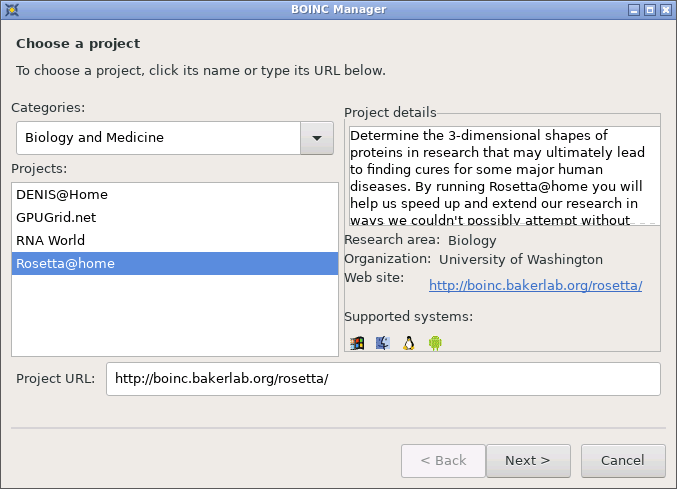 Adding Rosetta@home to your install
In order to receive computational tasks, you will need to sign up to the Rosetta project. This way your progress can be tracked by its systems. You can also view the progress of your tasks on the Rosetta website as well as your BOINC client.
Adding Rosetta@home to your install
In order to receive computational tasks, you will need to sign up to the Rosetta project. This way your progress can be tracked by its systems. You can also view the progress of your tasks on the Rosetta website as well as your BOINC client.
Once you have added the project, it will gather any updates and any tasks it needs to run. I would also highly recommend switching to the advanced view to better see the current task status: View -> Advanced View....
You can also set up any computing preferences so that BOINC does not overuse or block your CPU when you need it for other tasks. These values can be as high or low as you wish—anything is better than nothing!
In the Tasks pane, you are then able to see the current status of your tasks:
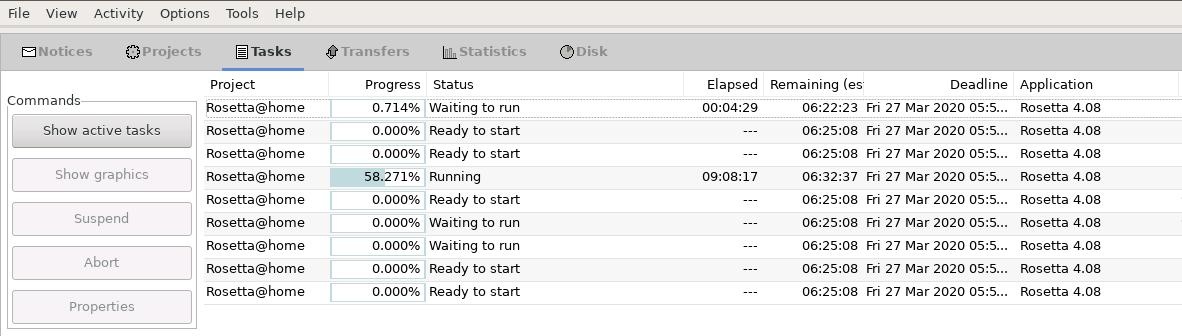 Task view of Rosetta@home
Task view of Rosetta@home
Folding@home
A similar solution to BOINC, operated by Stanford University:
Folding@home (FAH or F@h) is a distributed computing project for simulating protein dynamics, including the process of protein folding and the movements of proteins implicated in a variety of diseases. It brings together citizen scientists who volunteer to run simulations of protein dynamics on their personal computers. Insights from this data are helping scientists to better understand biology, and providing new opportunities for developing therapeutics.
Unlike BOINC, which allows you to add a multitude of projects, Folding@home has a singular purpose with a few different sub-types. In the case of COVID-19, the "Any" type prioritises this.
Installing
https://hub.docker.com/r/linuxserver/foldingathome
foldingathome:
image: linuxserver/foldingathome
container_name: foldingathome
environment:
- PUID=1000
- PGID=1000
- TZ=Europe/London
volumes:
- /path/to/data:/config
ports:
- 7396:7396
- 36330:36330 #optional
restart: unless-stoppedYou can access the web UI at http://<server_ip>:7396 (try accessing it via an InPrivate/Incognito window if you're having issues with your browser cache).
The built-in web server provides a very basic control to get you up and running (i.e. GPUs are only active when set to Medium or higher). For more fine grained control of individual devices, you can use the FAHControl app on a different device and connect remotely via port 36330. Bear in mind this is an unauthenticated connection.
If you wish to utilise your Nvidia GPU, you will need to install the container runtime provided by Nvidia on the host machine running the F@h container. Once nvidia-docker is installed on your host you will need to recreate the F@h container using the Nvidia container runtime instead (--runtime=nvidia) also adding an extra environment variable to enable the relevant devices for passthrough: -e NVIDIA_VISIBLE_DEVICES=all. This can also be set to a specific GPU's UUID, which can be discovered by running:
nvidia-smi --query-gpu=gpu_name,gpu_uuid --format=csv
Nvidia automatically mounts the GPU and drivers from your host into the Folding@home docker container. Our container has an environment variable set which ensures you are getting full use out of your GPU.
Don't forget to enter your user details into the web UI. When everything is set up and the work is downloaded from the servers, you should see the ongoing processes and the credits earned.
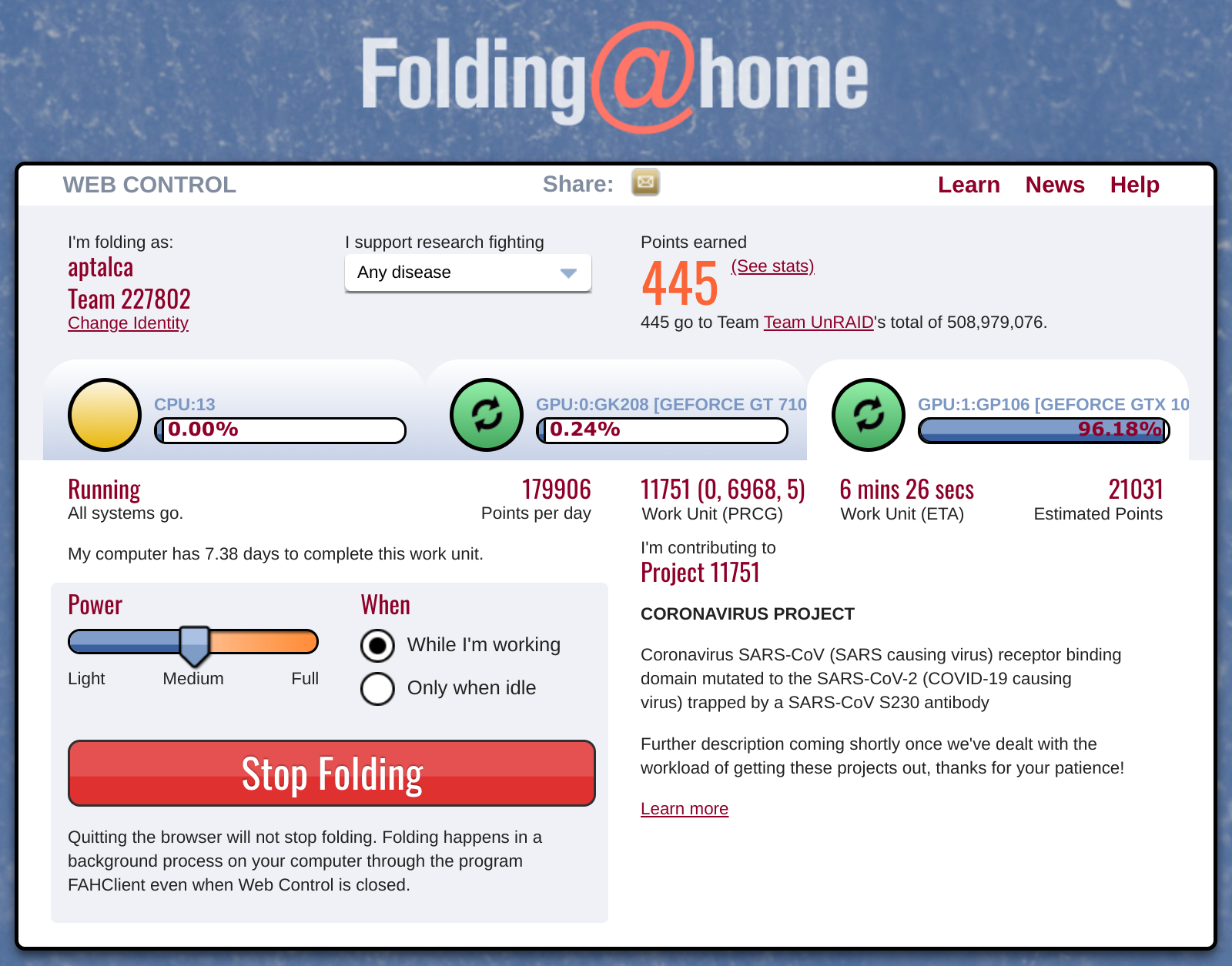 Folding@home built-in web UI
Now more than ever this is a time for everyone to come together to help prevent this kind of thing from happening again. It may seem innocuous now but doing something as simple as lending your CPU time for these projects does help.
Folding@home built-in web UI
Now more than ever this is a time for everyone to come together to help prevent this kind of thing from happening again. It may seem innocuous now but doing something as simple as lending your CPU time for these projects does help.
Happy folding!